|
受注:045-509-1970 |
技術サポート:[email protected] 平日9:00〜18:00 1営業日以内にご連絡を差し上げます |
化学情報
|
Synonyms | N/A | Storage (From the date of receipt) |
3 years -20°C powder 1 years -80°C in solvent |
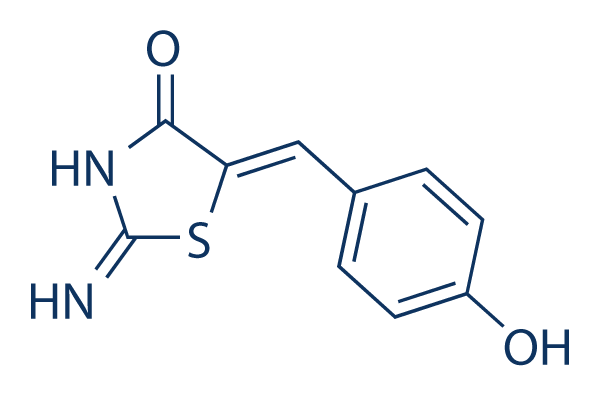
| 化学式 | C10H8N2O2S |
|||
| 分子量 | 220.25 | CAS No. | 1198097-97-0 | |
| Solubility (25°C)* | 体外 | DMSO | 44 mg/mL (199.77 mM) | |
| Water | Insoluble | |||
| Ethanol | Insoluble | |||
|
* <1 mg/ml means slightly soluble or insoluble. * Please note that Selleck tests the solubility of all compounds in-house, and the actual solubility may differ slightly from published values. This is normal and is due to slight batch-to-batch variations. |
||||
溶剤液(一定の濃度)を調合する
生物活性
| 製品説明 | Mirin is a potent Mre11–Rad50–Nbs1 (MRN) complex inhibitor, and inhibits Mre11-associated exonuclease activity. Mirin inhibits MRN-dependent activation of ATM. |
|---|---|
| in vitro | Mirin inhibits DSB-induced ATM activation, the ATM-dependent phosphorylation of the downstream targets Nbs1 and Chk2 and the MRN-dependent autophosphorylation of ATM at Ser1981 in response to DSBs. Mirin also inhibits the G2 checkpoint in TOSA4 cells, and homology-dependent DNA repair in HEK293 cells. [1] In cells with integrated HPV16 (SiHa), Mirin sensitizes HPV episomes to PA25 resulting in a ∼5-fold reduction of the PA25 IC50. [2] Pretreatment with mirin also decreases cell viability and inhibits proliferating cell nuclear antigen expression in cisplatin-treated human embryonic kidney 293 cells. [3] |
| in vivo | Mirin in nanoparticles resulted in a sharp impairment of tumor growth, associated with DDR activation, p53 accumulation, and cell death. |
プロトコル(参考用のみ)
| キナーゼアッセイ | Nuclease assay | |
|---|---|---|
| Reactions with oligonucleotide nonhairpin substrates contains 25 mM MOPS (pH 7.0), 60 mM KCl, 0.2% Tween 20, 2 mM DTT, 1 mM or 5 MnCl2 (or 5 mM MgCl2, or 5 mM CaCl2), 0.1 pmol of DNA substrate, and 0.3 pmol of Mre11 (or an equivalent amount of Mre11 complexed with Rad50) in a volume of 10 μl, and are incubated at 37°C for 30 min. SDS, EDTA, and proteinase K are then added to final concentrations of 0.2%, 5 mM, and 0.1 mg/ml, respectively, and incubated for another 15 min. 4 μl of each reaction is mixed with 4 μl of formamide loading buffer, and then loaded onto a sequencing gel containing 10% acrylamide and 7 M urea. After the run, each gel is analyzed using a phosphorimaging system. Reactions containing hairpin substrates are identical to those with nonhairpin substrates except that 3 pmol of Mre11 is added to reactions as indicated, and the reactions are incubated at room temperature overnight. Nonhomologous end-joining reactions contains 25 mM MOPS (pH 7.0), 60 mM KCl, 0.2% Tween 20, 2 mM DTT, 4 mM MgCl2, 2 mM MnCl2, 0.5 mM ATP, 4 ng of plasmid DNA, 10% polyethylene glycol, 0.01 pmol of human DNA ligase I, and 0.06 pmol of Mre11 or 0.1 units of E. coli exonuclease III (GIBCO-BRL), in a volume of 10 μl. After incubation at 37°C for 25 min, Tween 20 is added to a final concentration of 0.5%, and a 2.5 μl aliquot is amplified by PCR using primers DAR5 and DAR147. PCR products are cloned using the TA cloning kit and sequenced using an automated ABI Capillary Genetic Analyzer. | ||
| 細胞アッセイ | 細胞株 | HEK 293 cells |
| 濃度 | 100 μM | |
| 反応時間 | 24 h | |
| 実験の流れ | Human embryonic kidney (HEK) 293 cells are maintained in RPMI-1640 supplemented with 5% heat-inactivated fetal bovine serum, penicillin (100 U/mL), and streptomycin (100 mg/mL) in humidified air with 5% CO2 at 37 °C. Cells are given fresh medium at 48 h intervals. The cells are seeded in 96-well plates in regular growth medium. Cells are pretreated with mirin (100 μM for 1 h before the cisplatin (20 μM) treatment followed by incubation for 8 and 24 h. The MTT assay is performed using the EZ-Cytox cell viability assay kit according to the manufacturer's protocol and MTT reduction is measured at a 450 nm wavelength using a micro-plate reader. |
|
| 動物実験 | 動物モデル | Female BALB/c nude mice |
| 投薬量 | 50 mg/kg | |
| 投与方法 | ||
参考
カスタマーフィードバック
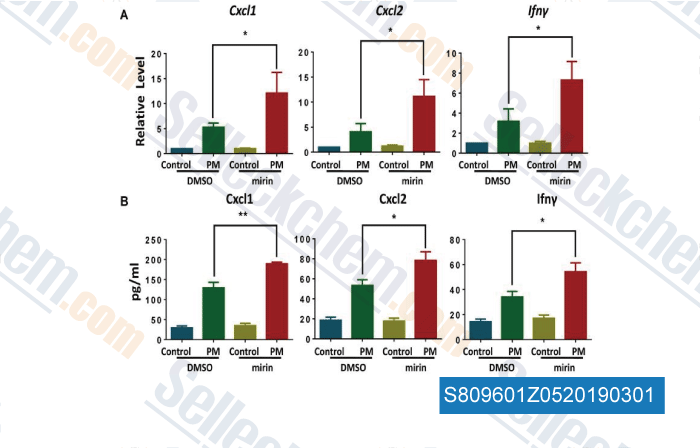
-
Data from [Data independently produced by , , Aging, 2018, 10(4):549-560]
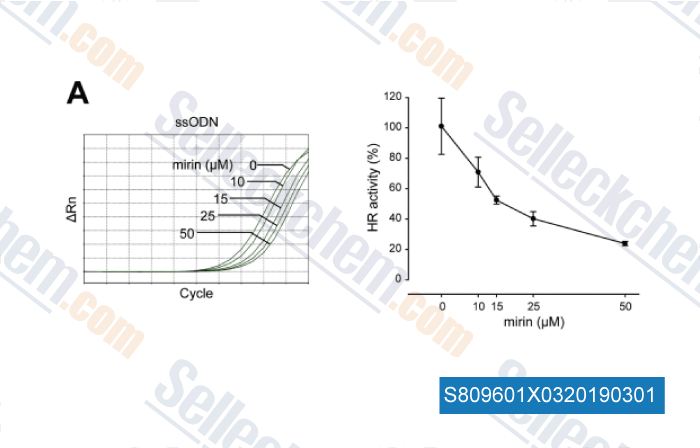
-
Data from [Data independently produced by , , DNA Repair, 2018, 70:67-71]
Selleckの高級品が、幾つかの出版された研究調査結果(以下を含む)で使われた:
| Replication fork uncoupling causes nascent strand degradation and fork reversal [ Nat Struct Mol Biol, 2023, 30(1):115-124] | PubMed: 36593312 |
| Replication fork uncoupling causes nascent strand degradation and fork reversal [ Nat Struct Mol Biol, 2023, 30(1):115-124] | PubMed: 36593312 |
| Multi-step processing of replication stress-derived nascent strand DNA gaps by MRE11 and EXO1 nucleases [ Nat Commun, 2023, 14(1):6265] | PubMed: 37805499 |
| Short-range end resection requires ATAD5-mediated PCNA unloading for faithful homologous recombination [ Nucleic Acids Res, 2023, 51(19):10519-10535] | PubMed: 37739427 |
| Short-range end resection requires ATAD5-mediated PCNA unloading for faithful homologous recombination [ Nucleic Acids Res, 2023, 10.1093/nar/gkad776] | PubMed: 37739427 |
| ATR protects ongoing and newly assembled DNA replication forks through distinct mechanisms [ Cell Rep, 2023, 42(7):112792] | PubMed: 37454295 |
| The MRN complex maintains the biliary-derived hepatocytes in liver regeneration through ATR-Chk1 pathway [ npj Regenerative Medicine, 2023, 20-2023)] | PubMed: None |
| The MRN complex maintains the biliary-derived hepatocytes in liver regeneration through ATR-Chk1 pathway [ NPJ Regen Med, 2023, 8(1):20] | PubMed: 37024481 |
| The KU-PARP14 axis differentially regulates DNA resection at stalled replication forks by MRE11 and EXO1 [ Nat Commun, 2022, 13(1):5063] | PubMed: 36030235 |
| ATM Regulates Differentiation of Myofibroblastic Cancer-Associated Fibroblasts and Can Be Targeted to Overcome Immunotherapy Resistance [ Cancer Res, 2022, 82(24):4571-4585.] | PubMed: 36353752 |
長期の保管のために-20°Cの下で製品を保ってください。
人間や獣医の診断であるか治療的な使用のためにでない。
各々の製品のための特定の保管と取扱い情報は、製品データシートの上で示されます。大部分のSelleck製品は、推薦された状況の下で安定です。製品は、推薦された保管温度と異なる温度で、時々出荷されます。長期の保管のために必要とされてそれと異なる温度で、多くの製品は、短期もので安定です。品質を維持するが、夜通しの積荷のために最も経済的な貯蔵状況を用いてあなたの送料を保存する状況の下に、製品が出荷されることを、我々は確実とします。製品の受領と同時に、製品データシートの上で貯蔵推薦に従ってください。
