- 阻害剤
- 研究分野別
- PI3K/Akt/mTOR
- Epigenetics
- Methylation
- Immunology & Inflammation
- Protein Tyrosine Kinase
- Angiogenesis
- Apoptosis
- Autophagy
- ER stress & UPR
- JAK/STAT
- MAPK
- Cytoskeletal Signaling
- Cell Cycle
- TGF-beta/Smad
- 化合物ライブラリー
- Popular Compound Libraries
- Customize Library
- Clinical and FDA-approved Related
- Bioactive Compound Libraries
- Inhibitor Related
- Natural Product Related
- Metabolism Related
- Cell Death Related
- By Signaling Pathway
- By Disease
- Anti-infection and Antiviral Related
- Neuronal and Immunology Related
- Fragment and Covalent Related
- Customize Library(compound antibody kit)
- FDA-approved Drug Library
- FDA-approved & Passed Phase I Drug Library
- Preclinical/Clinical Compound Library
- Bioactive Compound Library-I
- Bioactive Compound Library-II
- Kinase Inhibitor Library
- Express-Pick Library
- Natural Product Library
- Human Endogenous Metabolite Compound Library
- Alkaloid Compound LibraryNew
- Angiogenesis Related compound Library
- Anti-Aging Compound Library
- Anti-alzheimer Disease Compound Library
- Antibiotics compound Library
- Anti-cancer Compound Library
- Anti-cancer Compound Library-Ⅱ
- Anti-cancer Metabolism Compound Library
- Anti-Cardiovascular Disease Compound Library
- Anti-diabetic Compound Library
- Anti-infection Compound Library
- Antioxidant Compound Library
- Anti-parasitic Compound Library
- Antiviral Compound Library
- Apoptosis Compound Library
- Autophagy Compound Library
- Calcium Channel Blocker LibraryNew
- Cambridge Cancer Compound Library
- Carbohydrate Metabolism Compound LibraryNew
- Cell Cycle compound library
- CNS-Penetrant Compound Library
- Covalent Inhibitor Library
- Cytokine Inhibitor LibraryNew
- Cytoskeletal Signaling Pathway Compound Library
- DNA Damage/DNA Repair compound Library
- Drug-like Compound Library
- Endoplasmic Reticulum Stress Compound Library
- Epigenetics Compound Library
- Exosome Secretion Related Compound LibraryNew
- FDA-approved Anticancer Drug LibraryNew
- Ferroptosis Compound Library
- Flavonoid Compound Library
- Fragment Library
- Glutamine Metabolism Compound Library
- Glycolysis Compound Library
- GPCR Compound Library
- Gut Microbial Metabolite Library
- HIF-1 Signaling Pathway Compound Library
- Highly Selective Inhibitor Library
- Histone modification compound library
- HTS Library for Drug Discovery
- Human Hormone Related Compound LibraryNew
- Human Transcription Factor Compound LibraryNew
- Immunology/Inflammation Compound Library
- Inhibitor Library
- Ion Channel Ligand Library
- JAK/STAT compound library
- Lipid Metabolism Compound LibraryNew
- Macrocyclic Compound Library
- MAPK Inhibitor Library
- Medicine Food Homology Compound Library
- Metabolism Compound Library
- Methylation Compound Library
- Mouse Metabolite Compound LibraryNew
- Natural Organic Compound Library
- Neuronal Signaling Compound Library
- NF-κB Signaling Compound Library
- Nucleoside Analogue Library
- Obesity Compound Library
- Oxidative Stress Compound LibraryNew
- Phenotypic Screening Library
- PI3K/Akt Inhibitor Library
- Protease Inhibitor Library
- Protein-protein Interaction Inhibitor Library
- Pyroptosis Compound Library
- Small Molecule Immuno-Oncology Compound Library
- Mitochondria-Targeted Compound LibraryNew
- Stem Cell Differentiation Compound LibraryNew
- Stem Cell Signaling Compound Library
- Natural Phenol Compound LibraryNew
- Natural Terpenoid Compound LibraryNew
- TGF-beta/Smad compound library
- Traditional Chinese Medicine Library
- Tyrosine Kinase Inhibitor Library
- Ubiquitination Compound Library
-
Cherry Picking
You can personalize your library with chemicals from within Selleck's inventory. Build the right library for your research endeavors by choosing from compounds in all of our available libraries.
Please contact us at info@selleck.co.jp to customize your library.
You could select:
- 抗体
- 新製品
- お問い合わせ
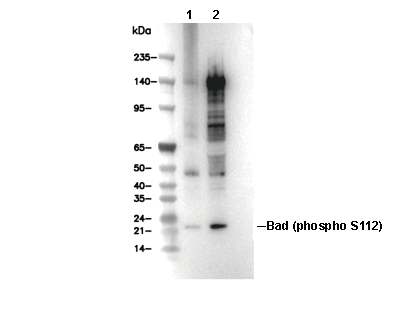
Phospho-Bad (Ser112) Antibody [F3L12]
Catalog No.: F3347
Application:
Reactivity:
当該製品は品切れ状态で、メールアドレスをご教示いただければ、お客様に返信いたします。
代表番号: 045-509-1970|電子メール:sales@selleck.co.jp
キーポイント
WB
転写条件(ウェット): 200 mA, 60 min,Recommended to use 0.22 μm PVDF 膜の使用をお勧めします。
転写条件(ウェット): 200 mA, 60 min,Recommended to use 0.22 μm PVDF 膜の使用をお勧めします。
使用情報
| Dilution |
|---|
|
| Application |
|---|
| WB, IP |
| Source |
|---|
| Rabbit Monoclonal Antibody |
| Reactivity |
|---|
| Rat, Human |
| Storage Buffer |
|---|
| PBS, pH 7.2+50% Glycerol+0.05% BSA+0.01% NaN3 |
| Storage (from the date of receipt) |
|---|
| -20°C (avoid freeze-thaw cycles), 2 years |
| Predicted MW Observed MW |
|---|
| 18 kDa 23 kDa |
| *なぜ予測分子量と実際の分子量が異なるのか? 下記の原因により、実際の分子量が予測と異なる:タンパク質の翻訳後修飾(リン酸化/糖鎖付加),スプライシングバリアント,イソフォーム,相対的な電荷,ポリマー。 |
| ポジティブコントロール | HeLa cells (Calyculin A, 100 ng/ml, 30 min); C6 cells (Calcyculin A treated) |
|---|---|
| ネガティブコントロール | C6 cells; HeLa cells |
プロトコール
| WB |
|---|
Experimental Protocol:
Sample preparation
1. Tissue: Lyse the tissue sample by adding an appropriate volume of ice-cold RIPA/NP-40 Lysis Buffer (containing Protease Inhibitor Cocktail, Phosphatase Inhibitor Cocktail),and homogenize the tissue at a low temperature or lyse it by sonication on ice, then incubate on ice for 30 minutes. 2. Adherent cell: Aspirate the culture medium and wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/NP-40 Lysis Buffer (containing Protease Inhibitor Cocktail, Phosphatase Inhibitor Cocktail), sonicate to lyse the cells, and incubate on ice for 30 minutes. 3. Suspension cell: Transfer the culture medium to a pre-cooled centrifuge tube. Centrifuge and aspirate the supernatant. Wash the cells with ice-cold PBS twice. Lyse the cells by adding an appropriate volume of RIPA/NP-40 Lysis Buffer (containing Protease Inhibitor Cocktail, Phosphatase Inhibitor Cocktail), sonicate to lyse the cells, and incubate on ice for 30 minutes. 4. Place the lysate into a pre-cooled microcentrifuge tube. Centrifuge at 4°C for 15 min. Collect the supernatant;
5. Remove a small volume of lysate to determine the protein concentration;
6. Combine the lysate with protein loading buffer. Boil 20 µL sample under 95-100°C for 5 min. Centrifuge for 5 min after cool down on ice.
Electrophoretic separation
1. According to the concentration of extracted protein, load appropriate amount of protein sample and marker onto SDS-PAGE gels for electrophoresis. Recommended separating gel (lower gel) concentration: 10%. Reference Table for Selecting SDS-PAGE Separation Gel Concentrations 2. Power up 80V for 30 minutes. Then the power supply is adjusted (110 V~150 V), the Marker is observed, and the electrophoresis can be stopped when the indicator band of the predyed protein Marker where the protein is located is properly separated. (Note that the current should not be too large when electrophoresis, too large current (more than 150 mA) will cause the temperature to rise, affecting the result of running glue. If high currents cannot be avoided, an ice bath can be used to cool the bath.)
Transfer membrane
1. Take out the converter, soak the clip and consumables in the pre-cooled converter;
2. Activate PVDF membrane with methanol for 1 min and rinse with transfer buffer;
3. Install it in the order of "black edge of clip - sponge - filter paper - filter paper - glue -PVDF membrane - filter paper - filter paper - sponge - white edge of clip"; 4. The protein was electrotransferred to PVDF membrane. ( 0.22 µm PVDF membrane is recommended )) Reference Table for Selecting PVDF Membrane Pore Size Specifications Recommended conditions for wet transfer: 200 mA, 60 min. ( Note that the transfer conditions can be adjusted according to the protein size. For high-molecular-weight proteins, a higher current and longer transfer time are recommended. However, ensure that the transfer tank remains at a low temperature to prevent gel melting.)
Block
1. After electrotransfer, wash the film with TBST at room temperature for 5 minutes;
2. Incubate the film in the blocking solution ( recommending 5% BSA solution)
for 1 hour at room temperature;
3. Wash the film with TBST for 3 times, 5 minutes each time.
Antibody incubation
1. Use 5% skim milk powder to prepare the primary antibody working liquid (recommended dilution ratio for primary antibody 1:5000), gently shake and incubate with the film at 4°C overnight; 2. Wash the film with TBST 3 times, 5 minutes each time;
3. Add the secondary antibody to the blocking solution and incubate with the film gently at room temperature for 1 hour;
4. After incubation, wash the film with TBST 3 times for 5 minutes each time.
Antibody staining
1. Add the prepared ECL luminescent substrate (or select other color developing substrate according to the second antibody) and mix evenly;
2. Incubate with the film for 1 minute, remove excess substrate (keep the film moist), wrap with plastic film, and expose in the imaging system. |
生物学的記述
| Specificity |
|---|
| Phospho-Bad (Ser112) Antibody [F3L12] detects endogenous levels of total Bad protein only when it is phosphorylated at Ser112. |
| タンパク質の局在 |
|---|
| 細胞質、細胞内膜系、ミトコンドリア、ミトコンドリア外膜 |
| Uniprot ID |
|---|
| Q92934 |
| Clone |
|---|
| F3L12 |
| Synonym(s) |
|---|
| BBC6; BCL2L8; BAD; Bcl2-associated agonist of cell death; Bcl-2-binding component 6; Bcl-2-like protein 8; Bcl-xL/Bcl-2-associated death promoter; Bcl2 antagonist of cell death; Bcl2-L-8 |
| Background |
|---|
| Phospho-Bad Ser112 refers to the serine 112 phosphorylation site on Bad, a 184-amino acid proapoptotic BH3-only member of the Bcl-2 family that forms heterodimers with anti-apoptotic Bcl-2 and Bcl-xL through its alpha-helical BH3 domain spanning residues 103 to 127, with key residues Leu107, Asp112, and Leu118 crucial for insertion into the Bcl-xL binding groove and displacement of Bax or Bak to trigger mitochondrial outer membrane permeabilization and apoptosis. Ser112 is located N-terminal to the BH3 amphipathic helix in a flexible loop, and phosphorylation at this site induces conformational changes that expose 14-3-3 phospho-binding motifs, specifically the mode I RSXpSXP motif at residues 112 to 117 and, when Ser136 is also phosphorylated, the mode II RXYpSXP motif at residues 136 to 141. This modification disrupts the amphipathic nature of the BH3 helix and inhibits docking to Bcl-xL, instead promoting cytosolic sequestration of Bad by 14-3-3 proteins. Ser112 phosphorylation mediates survival signaling in response to growth factors such as IL-3 or EGF, which activate the Ras/Raf to MAPK/ERK to p90RSK pathway or mitochondria-anchored PKA, leading to Bad phosphorylation and sequestration in 14-3-3 complexes, thereby blocking its pro-apoptotic activity and supporting cell survival and proliferation. Dephosphorylation by PP2A, particularly upon withdrawal of survival signals, destabilizes 14-3-3 binding and, together with loss of phosphorylation at Ser136 (by Akt) and Ser155 (by PKA), releases Bad to antagonize Bcl-xL or Bcl-2 and promote apoptosis. This regulatory mechanism links cytokine and PI3K-independent MAPK pathways to the mitochondrial apoptosis rheostat and is especially important for lymphocyte and growth factor-dependent cell survival. Hyperphosphorylated Bad mediated by oncogenic Ras or MAPK pathways contributes to therapy resistance in cancers such as leukemia and carcinomas, while reactivation of PP2A can resensitize tumors to apoptosis. |
| References |
|---|
技術サポート
ストックの作り方、阻害剤の保管方法、細胞実験や動物実験の際に注意すべき点など、製品を取扱う時に問い合わせが多かった質問に対しては取扱説明書でお答えしています。
他に質問がある場合は、お気軽にお問い合わせください。
* 必須